Conservation and Evolutionary Genetics Group |
||||||||||
| HOME | PEOPLE | PROJECTS | PUBLICATIONS | RESOURCES | COURSES | NETWORKS | OUTREACH | CONTACT | ||
|
Late Pleistocene and Holocene Population Changes and Extinctions |
One of the greatest uncertainties regarding the biotic impact of anticipated global warming is the influence of climatic change on the abundance of different genotypes and on their geographic distribution. Questions of biotic response to global warming are critical in the Arctic, as polar regions are particularly sensitive to climate change and are expected to experience continued warming and consequent displacements of species' ranges and biomes. The Pleistocene climatic and environmental history of the Holarctic has been turbulent. Understanding mammalian responses to past climatic changes is important to predicting and potentially mitigating the impact of future climatic changes on extant species. We investigate relationships between genetic and environmental change during the late Pleistocene and Holocene for a variety of species, and also study how they are related to living populations and species to better understand their evolution.

Image: George Teichmann
Reconstructions of past environments and climates have generally used plant and animal macrofossil, palynological, paleolimnological and ice-core data. However, these approaches provide little or no information on genotypic changes that might have occurred during past periods of climatic change. In addition, continuous plant macrofossil, palynological or paleolimnological records extending through the last glacial cycle remain sparse and sometimes difficult to interpret. Recent ancient DNA studies have shown that brown bears (Ursus arctos) in Beringia experienced population turnovers in the period from 60 to 14 thousand years before present (ka BP), implying habitat changes and a genotypic response not revealed by other analyses. The causes for these Quaternary population turnovers are unclear. These studies demonstrate the great potential of ancient DNA analysis to illuminate the impact of environmental change on the diversity, geographic structure and ultimately, the fate of mammalian populations.
 |
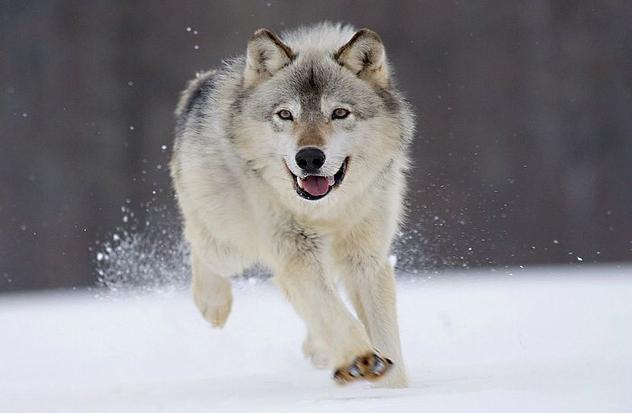
We explore the generality of these past turnover events by analyzing patterns of mitochondrial DNA sequence change over the past 40,000 years in more large mammal species with different ecological requirements. Because the habitat preferences differ, we might expect them to show unique responses to changes in late Quaternary climate and vegetation. DNA sequenced from bones of these species will provide a record of genetic variability and genetic turnover reflecting changes in population size and the frequency of extinction and recolonization events. |
We are currently working with extinct camels and caribou from North America, and horses, canids, bovids and lagomorphs from Europe.
 |
 |
Jennifer Leonard is leading this research line, Carlos Domínguez Sarabia is working on it as PhD student and Anna Cornellas as lab technician.
Related publications
Peer-reviewed Articles
|
Evolutionary history of saber-toothed cats based on ancient mitogenomics
| ||||
|
|
|||
|
The role of canids in ritual and domestic contexts: new ancient DNA insights from complex hunter-gatherer sites in prehistoric Central California
|
||||
|
Ancient DNA analysis affirms the canid from Altai as a primitive dog
|
||||
|
Burying dogs in ancient cis-Baikal, Siberia: Temporal trends and relationships with human diet and subsistence practices
|
||||
|
Discovery of lost diversity of paternal horse lineages using ancient DNA
|
||||
|
Species-specific responses of Late Quaternary megafauna to climate and humans
|
||||
|
Nuclear copies of mitochondrial genes: another problem for ancient DNA
|
Funding
 |
|
© 2021 CONSEVOL |


